Medical image analysis has changed a lot in the last decade thanks to deep learning. However, taking these models from a research paper to a tool that a radiologist or a researcher can actually use every day is still difficult: you need to install dependencies, manage GPUs, replicate pre and post-processing pipelines… In this post I explore TOMAAT, an open-source framework that proposes an elegant solution: offering these models as a cloud service.
What is TOMAAT?
TOMAAT is a framework presented by Milletari et al. in 2018. It allows researchers to deploy deep learning models for volumetric medical image analysis (MRI, CT, PET, 3D ultrasound…) as services that can be accessed from the cloud. The idea is simple but powerful: the researcher who trains the model “wraps” it in a server, and any user can connect to it to get predictions without installing anything locally.
Instead of asking the user to reproduce the whole execution environment, TOMAAT takes the model to where the computing power is and only sends the results back.
Architecture: three pieces that fit together
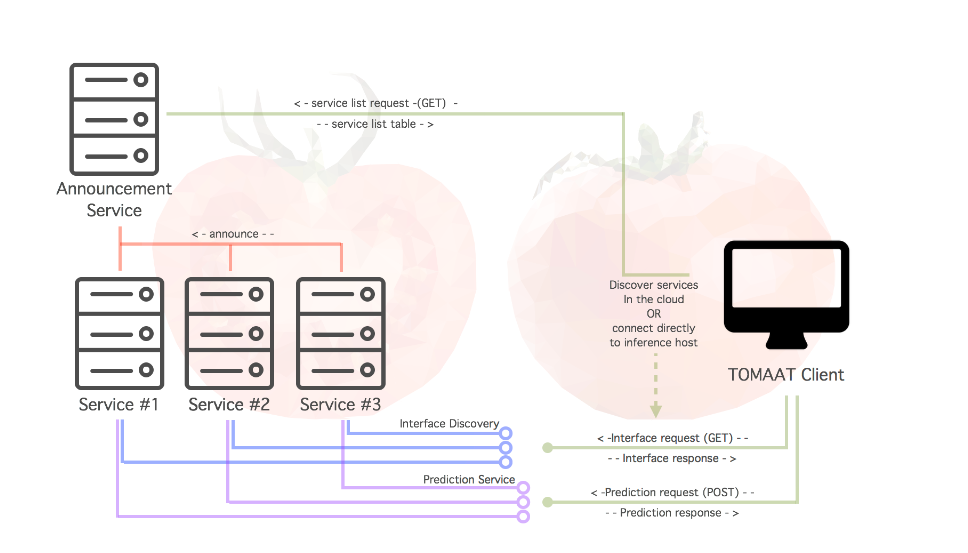
The framework has three main components:
- Server — wraps the trained model together with its pre and post-processing pipelines, and exposes it on the cloud.
- Client — communicates with the server to send the user’s data and receive the predictions.
- Service registry — a directory that tells clients which endpoints (models) are available in the cloud.
This separation is very common in cloud-native architectures and offers several advantages:
| Advantage | Why it matters |
|---|---|
| Scalability | More servers can be added when demand grows |
| Reproducibility | The environment is fixed on the server, not on the client |
| Accessibility | Any lab can use the model without owning a GPU |
| Collaboration | Researchers share models as services, not as code |
Why is it interesting from a cloud point of view?
TOMAAT is a good example of how to apply cloud computing concepts to a specific scientific problem. Instead of treating the cloud as just “a bigger remote computer”, it takes advantage of the Software as a Service (SaaS) model: the user consumes a feature (segmenting an organ, detecting a lesion) without worrying about the infrastructure behind it.
A simple example of how a client could call one of these services would look like this:
import requests
# Send a 3D volume to the model endpoint
with open("patient_001.nii.gz", "rb") as f:
response = requests.post(
"https://tomaat-registry.org/segmentation/prostate",
files={"volume": f}
)
segmentation = response.content # segmented volume comes back
The user does not know (and does not need to know) whether there is a GPU, a cluster, or several Docker containers running with Kubernetes behind the scenes.
Limitations and reflection
Not everything is perfect. Sending medical images to an external server raises important challenges in privacy and legal compliance (GDPR in Europe, HIPAA in the US). Network latency can also be a problem when working with large 3D volumes. For these reasons, many hospitals choose to deploy this kind of framework on private clouds instead of public ones.
Even so, TOMAAT shows an important idea: the future of medical image analysis will probably not be each lab training and running its own models, but an ecosystem of interoperable services accessible from the cloud.
Conclusion
Beyond the specific case of TOMAAT, the interesting point is to see how cloud computing principles —scalability, accessibility, separation between infrastructure and application— can be applied to a field as specific as medical imaging. And how, by doing this, the gap between research and clinical practice becomes smaller.
References
- Milletari, F., Frei, J., Aboulatta, M., Vivar, G., & Ahmadi, S. A. (2018). TOMAAT: volumetric medical image analysis as a cloud service. arXiv preprint arXiv:1803.06784.
- *Figure 1. TOMAAT architecture overview. Source: Milletari et al. (2018).